OpenMM
Computing Applications (HPC) MIT (Free)
About
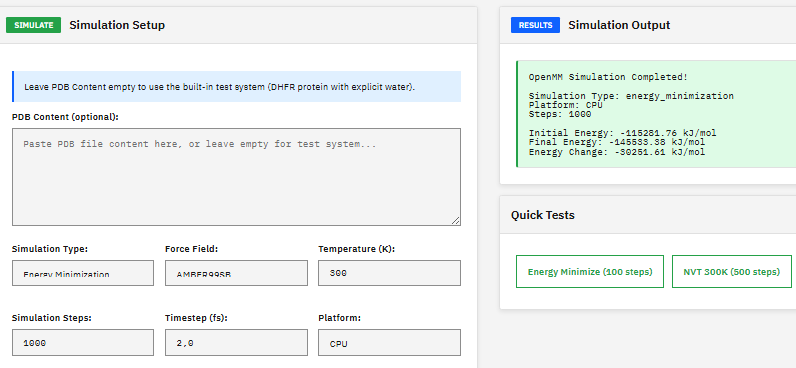
A molecular dynamics toolkit for simulating biomolecules, providing energy minimization, NVT/NPT ensemble sampling, and support for AMBER and CHARMM force fields with GPU-accelerated calculations via CUDA and OpenCL.
Key Features
- Force fields: AMBER, CHARMM, OPLS
- Ensembles: NVT, NPT, NVE
- GPU: CUDA, OpenCL acceleration
- Integrators: Langevin, Verlet, Brownian
Citation
Eastman, P. et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLOS Comput. Biol. 13, e1005659 (2017). DOI:10.1371/journal.pcbi.1005659
