OpenMM
Computing Applications (HPC) MIT (Free)
About
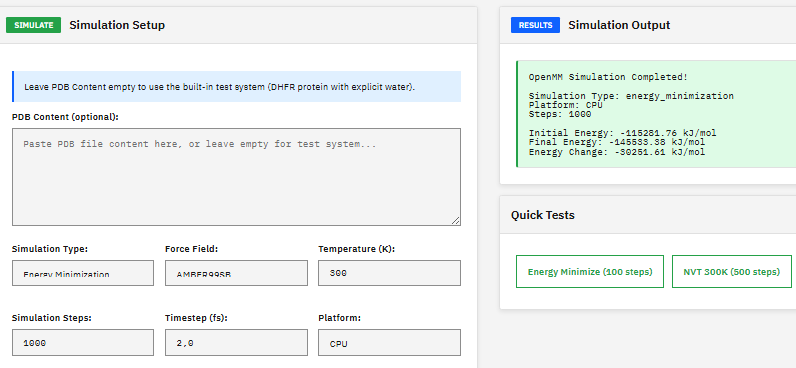
A molecular dynamics toolkit for simulating biomolecules, providing energy minimization, NVT/NPT ensemble sampling, and support for AMBER and CHARMM force fields with GPU-accelerated calculations via CUDA and OpenCL.
Key Features
- Force fields: AMBER, CHARMM, OPLS
- Ensembles: NVT, NPT, NVE
- GPU: CUDA, OpenCL acceleration
- Integrators: Langevin, Verlet, Brownian
Citation
Eastman, P. et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLOS Comput. Biol. 13, e1005659 (2017). DOI:10.1371/journal.pcbi.1005659
Frequently Asked Questions
What is OpenMM?
OpenMM is a computing applications (hpc) application available in the Paramus App Store. A molecular dynamics toolkit for simulating biomolecules, providing energy minimization, NVT/NPT ensemble sampling, and support for AMBER and CHARMM force fields with GPU-accelerated calculations via CUDA and OpenCL.
Is OpenMM free to use?
Yes. OpenMM is distributed under the MIT (Free) license and is available at no cost through the Paramus App Store.
How do I install OpenMM?
OpenMM is installed through Paramus Chemistry OS, an on-premise Windows platform for computational chemistry. Open the Paramus App Store in your local installation and select OpenMM for one-click deployment.
What are the key features of OpenMM?
Key features of OpenMM include: Force fields: AMBER, CHARMM, OPLS; Ensembles: NVT, NPT, NVE; GPU: CUDA, OpenCL acceleration; Integrators: Langevin, Verlet, Brownian.
What type of application is OpenMM?
OpenMM belongs to the “Computing Applications (HPC)” category in the Paramus App Store. It runs on Paramus Chemistry OS and can also be accessed through Paramus Cloud for supported workflows.
What platform does OpenMM run on?
OpenMM runs on Paramus Chemistry OS, a Windows-based on-premise platform that provides local compute power for demanding simulations. It requires a Paramus OS installation with appropriate hardware resources.
Can OpenMM be automated or integrated with AI workflows?
Yes. OpenMM is available as part of the Paramus ecosystem which supports MCP (Model Context Protocol) tools for AI-driven automation. This enables integration with large language models and automated research pipelines.
How should I cite OpenMM in publications?
The recommended citation for OpenMM is: Eastman, P. et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLOS Comput. Biol. 13, e1005659 (2017). DOI:10.1371/journal.pcbi.1005659
